
Population-Specific Genomic Data Improves Disease Risk Prediction in the Taiwanese Han Population
Recognizing this gap, researchers established HiGenome, a comprehensive genomic and longitudinal health database focused on the Taiwanese Han population. By integrating genetic sequencing data with detailed medical records from more than 323,000 participants, the team created one of the most extensive ancestry-specific datasets in East Asia. This integration allowed them not only to identify genetic variants associated with disease, but also to observe how those risks unfold over time in real-world clinical settings.
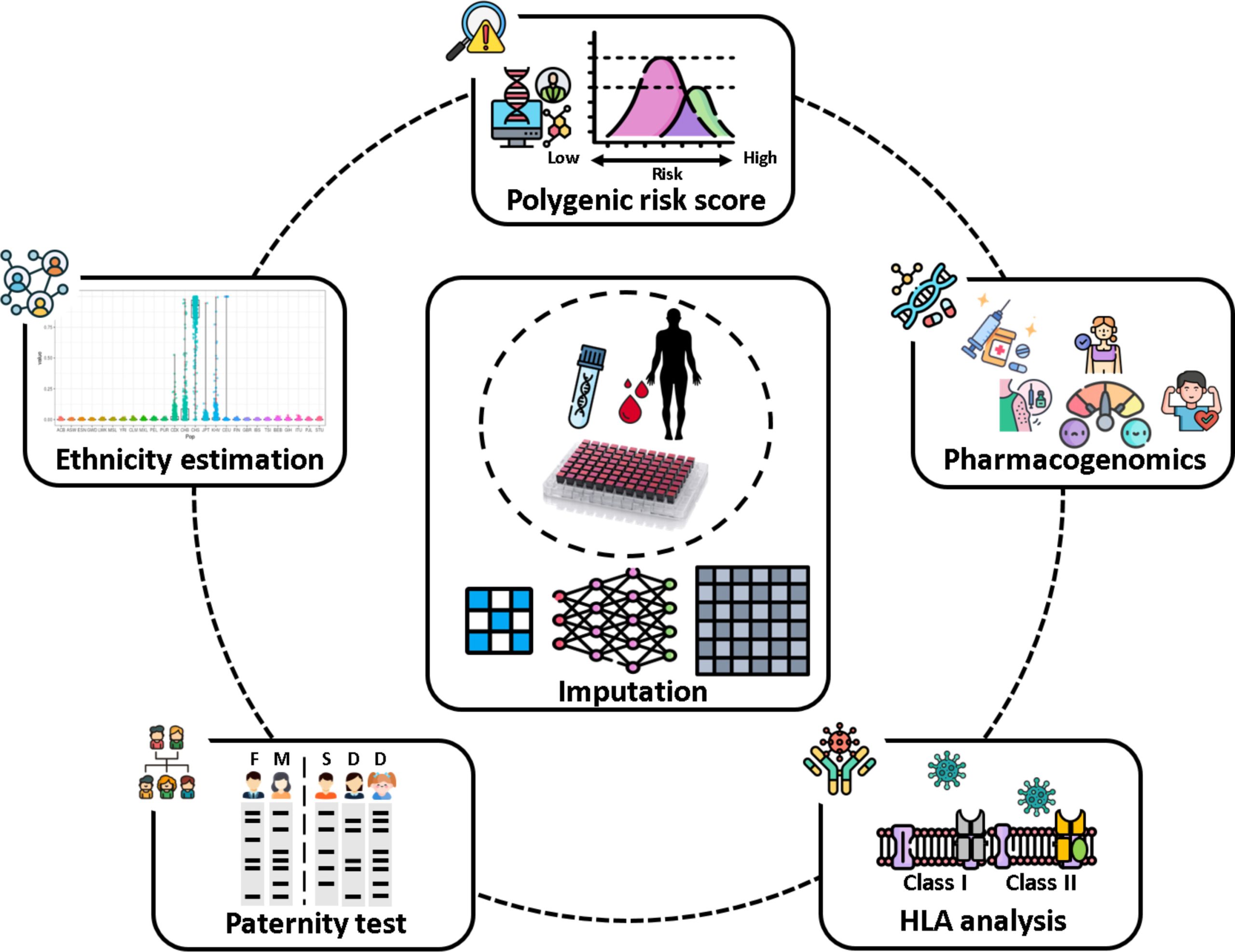
Using genome-wide association studies across 1,085 quantitative traits and diseases, the researchers mapped patterns of genetic influence spanning metabolic disorders, cardiovascular conditions, autoimmune diseases, and more. They also conducted phenome-wide analyses to examine how specific genetic variants were linked to multiple clinical outcomes. One key outcome of the project was the development of polygenic risk scores (PRS) for 238 diseases. These scores combine the small effects of numerous genetic variants to estimate an individual’s overall genetic susceptibility.
Importantly, the study revealed population-specific genetic associations that differed in strength or prevalence compared to those observed in cohorts. In some cases, variants that appeared modest in global datasets showed stronger predictive value within the Taiwanese Han population. By training polygenic risk models directly on ancestry-matched data, predictive accuracy improved significantly for certain conditions, including metabolic and kidney-related diseases.
The inclusion of longitudinal health records added another layer of insight. Rather than offering static snapshots of genetic risk, the database allowed researchers to observe how inherited susceptibility interacted with age, lifestyle, and clinical progression. This time-based perspective strengthens the potential for early intervention strategies tailored to individuals at elevated risk.
What makes this work distinctive is not only its scale, but its emphasis on representation. Precision medicine depends on understanding biological variation across populations. Without inclusive datasets, healthcare systems risk widening disparities rather than reducing them.
As genetic testing becomes more integrated into routine healthcare, tools built on diverse, population-specific data will become increasingly important. Resources such as HiGenome may support earlier detection, targeted prevention strategies, and more informed clinical decisions for millions of individuals. In the near future, ancestry-aware genomic research could help ensure that advances in precision medicine benefit global populations equitably, rather than privileging only those already well represented in research databases.
Reference
T.-Y. Liu et al., “Diversity and longitudinal records: Genetic architecture of disease associations and polygenic risk in the Taiwanese Han population,“ Science Advances, 2025, doi:10.1126/sciadv.adt0539.

Publication Title: Diversity and longitudinal records: Genetic architecture of disease associations and polygenic risk in the Taiwanese Han population
Journal Title: Science Advances
Publisher: AAAS
Year: 2025
Subject: Genome
Research Footprints:
HiGenome; polygenic risk score
